Decision trees - Classification trees
Classification Trees
Very similar to regression trees except it is used to predict a qualitative response rather than a quantitative one
We predict that each observation belongs to the most commonly occuring class of the training observations in a given region
Fitting classification trees
We use recursive binary splitting to grow the tree
Instead of RSS, we can use:
Gini index: \(G = \sum_{k=1}^K \hat{p}_{mk}(1-\hat{p}_{mk})\)
This is a measure of total variance across the \(K\) classes. If all of the \(\hat{p}_{mk}\) values are close to zero or one, this will be small
The Gini index is a measure of node purity small values indicate that node contains predominantly observations from a single class
In
R, this can be estimated using thegain_capture()function.
Classification tree - Heart Disease Example
- Classifying whether 303 patients have heart disease based on 13 predictors (
Age,Sex,Chol, etc)
1. Split the data into a cross-validation set
How many folds do I have?
2. Create a model specification that tunes based on complexity, \(\alpha\)
3. Fit the model on the cross validation set
What \(\alpha\)s am I trying?
5. Choose \(\alpha\) that minimizes the Gini Index
6. Fit the final model
7. Examine how the final model does on the full sample
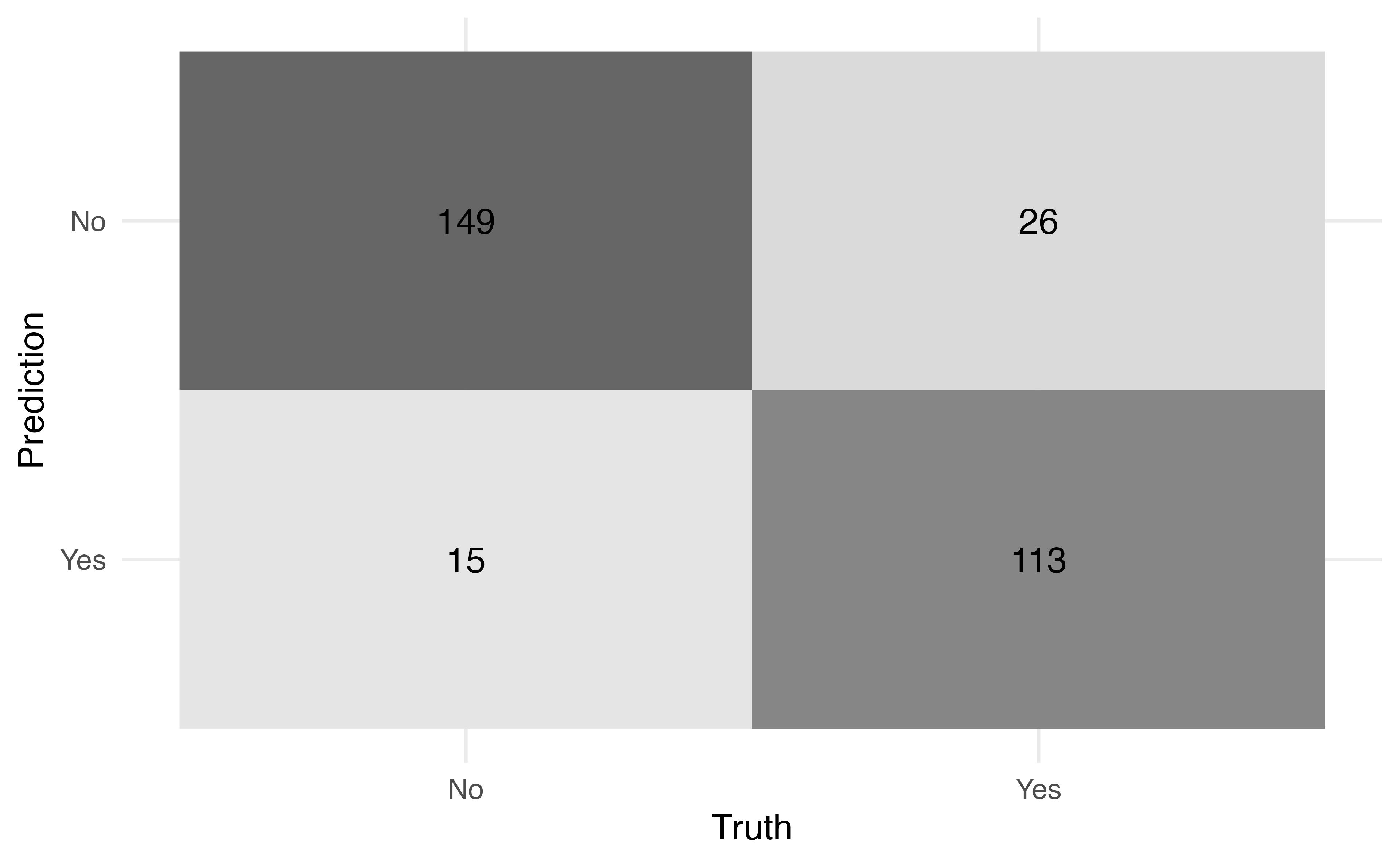
Decision trees
Pros
- simple
- easy to interpret
Cons
- not often competitive in terms of predictive accuracy
- Next we will discuss how to combine multiple trees to improve accuracy
Dr. Lucy D’Agostino McGowan adapted from slides by Hastie & Tibshirani